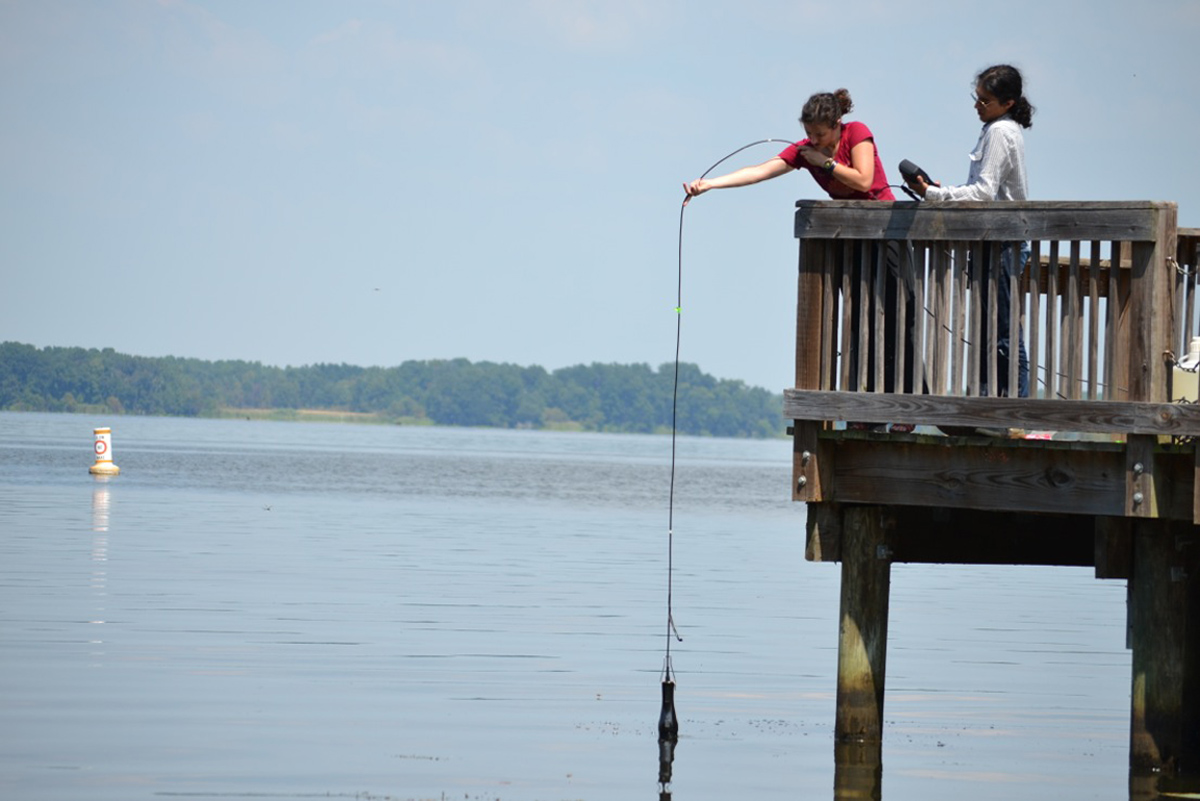
 Georgia Tech graduate students Despina Tsementzi and Miguel Rodriguez prepare to sample water from Lake Seminole in Georgia, one of the lakes on the Chattahoochee River system. The water samples will be used to study microbial communities as part of research in the laboratory of Kostas Konstantinidis at Georgia Tech. (Photo Courtesy of K. Konstantinidis and D. Tsementzi.) Georgia Tech graduate students Despina Tsementzi and Miguel Rodriguez prepare to sample water from Lake Seminole in Georgia, one of the lakes on the Chattahoochee River system. The water samples will be used to study microbial communities as part of research in the laboratory of Kostas Konstantinidis at Georgia Tech. (Photo Courtesy of K. Konstantinidis and D. Tsementzi.) |
By John Toon, Georgia Tech Research News
Microbes of interest to clinicians and environmental scientists rarely exist in isolation. Organisms essential to breaking down pollutants or causing illness live in complex communities, and separating one microbe from hundreds of companion species can be challenging for researchers seeking to understand environmental issues or disease processes.
A new National Science Foundation-supported project will provide computational tools designed to help identify and characterize the gene diversity of the residents of these microbial communities. The project, being done by researchers at the Georgia Institute of Technology and Michigan State University, will allow clinicians and scientists to compare the genomic information of organisms they encounter against the growing volumes of data provided by the world’s scientific community.
The tools will be hosted on a web server designed to be used by researchers who may not have training in the latest bioinformatics techniques. A prototype system containing a limited number of computational tools is already available at http://enve-omics.ce.gatech.edu and is attracting more than 500 users each month.
“Across many areas of science, we are dealing with communities of microorganisms, and one challenge we’ve had is to identify them because we haven’t had good tools to tell apart individual microbes from the mixtures,” said Kostas Konstantinidis, an associate professor in the School of Civil and Environmental Engineering at Georgia Tech and the project’s principal investigator. “Our tools will be designed to deal with the genomes of whole communities of organisms.”
Current techniques identify individual microbes by examining their small subunit ribosomal RNA (SSU rRNA) genes, but the new tools will allow scientists to analyze entire genomes and meta-genomes.
“With the dawn of the genomic era, we can now get the whole genome of these organisms to see not only the ribosomal RNA, but also all the genes in the genome to get a better understanding of what the each organism’s potential might be,” said Konstantinidis. “There will be many advantages for looking at all the genes instead of just one, the SSU rRNA, such as to identify which organisms encode toxins or the enzymes for breaking down pollutants.”
Collaborators on the three-year project include scientists who operate the Ribosomal Database Project at Michigan State University: Jim Tiedje, director of Michigan State University’s Center for Microbial Ecology and James Cole, a Michigan State University research assistant professor and director of the Ribosomal Database Project.
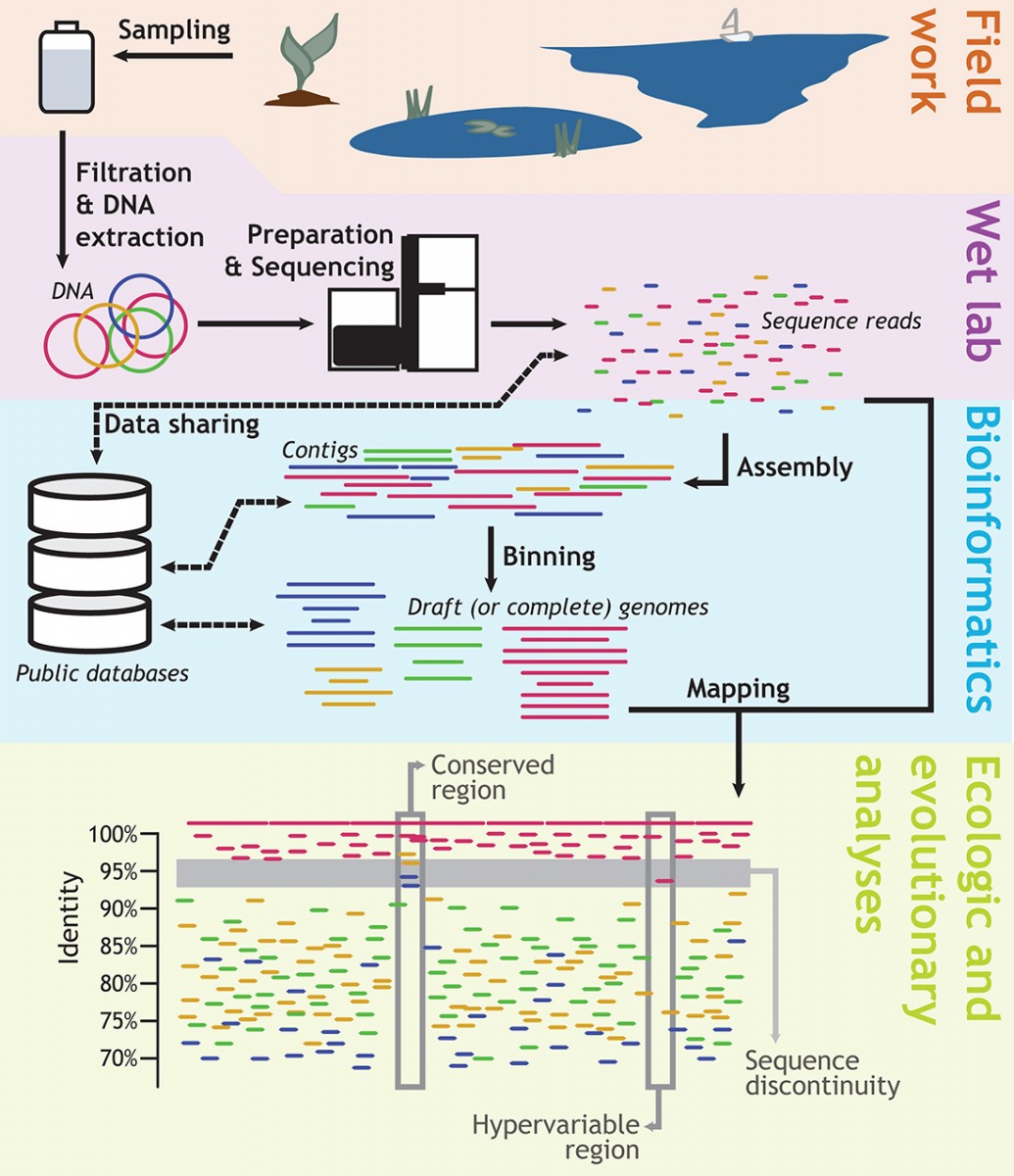 Diagram shows the major steps taken in research conducted on communities of microbes in lake water by the laboratory of Kostas Konstantinidis at Georgia Tech. The focus of the NSF-supported project will be to develop the bioinformatics tools that will enable genomic and meta-genomic research. (Modified with permission from Microbe Magazine, March 2014, p. 111) Diagram shows the major steps taken in research conducted on communities of microbes in lake water by the laboratory of Kostas Konstantinidis at Georgia Tech. The focus of the NSF-supported project will be to develop the bioinformatics tools that will enable genomic and meta-genomic research. (Modified with permission from Microbe Magazine, March 2014, p. 111) |
The ability to identify and enumerate the organisms in complex communities using culture-independent, genomic technologies and associated bioinformatics algorithms is becoming more important as scientists study organisms that can’t be grown in the lab. The majority of the world’s organisms resist traditional lab culture, meaning they have to be studied in the field and identified through genetic information.
Konstantinidis and his research group are studying such communities in the water of lakes in Chattahoochee River system in Georgia and elsewhere. They are examining how these communities respond to perturbations, such as oil or pesticide spills, and the role that different members of the community play in breaking down pollutants.
“These tools actually come from our research practice,” said Konstantinidis. “We came to the point where we couldn’t process the data to answer the questions we wanted to ask. That led us to this new project to develop the tools we and others need to interrogate the data and get the information we are looking for.”
A single liter of lake water may contain as many as 500 different species, and together, their genomic information can total tens of billions of gene-coding letters. From Lake Lanier alone, the team has generated 200 gigabytes of genomic data.
“We want to figure out what organisms are there, and what genes they encode,” Konstantinidis explained. “The tools we are developing will allow us to do this.”
The tools developed in the project will be useful to both clinical microbiologists and environmental researchers. “This will not be specific to any one discipline,” he said. “As long as people are working with microbes, this will be helpful to them because some of the questions are universal.”
The system will also be built to provide user-friendly help to scientists who may not have training in the latest genomic and bioinformatics techniques. “There is a big need for big data analysis, and there are not many trained people right now,” Konstantinidis said. “These tools will make the lives of researchers easier.”
Among the challenges ahead is building an infrastructure able to handle the growing amounts of genomic information produced worldwide.
“We will have to develop some computational solutions for the problems of keeping up with all the new data becoming available,” said Konstantinidis. “We need to make tools that have high throughput to keep up with data volumes that are increasing geometrically.”
The system will initially operate on servers at Georgia Tech and Michigan State University, but if demand and data grow, additional resources may be sought, such as the National Science Foundation’s XSEDE supercomputer.
This research is supported by the National Science Foundation under award DBI-1356288. The opinions expressed in this article are those of the authors and do not necessarily reflect the official views of the National Science Foundation.
Kostas Konstantinidis is the Carlton S. Wilder Junior Faculty Professor in the Georgia Tech School of Civil and Environmental Engineering.
